DREAMS: deep read-level error model for sequencing data applied to low-frequency variant calling and circulating tumor DNA detection, Genome Biology
Por um escritor misterioso
Last updated 30 julho 2024
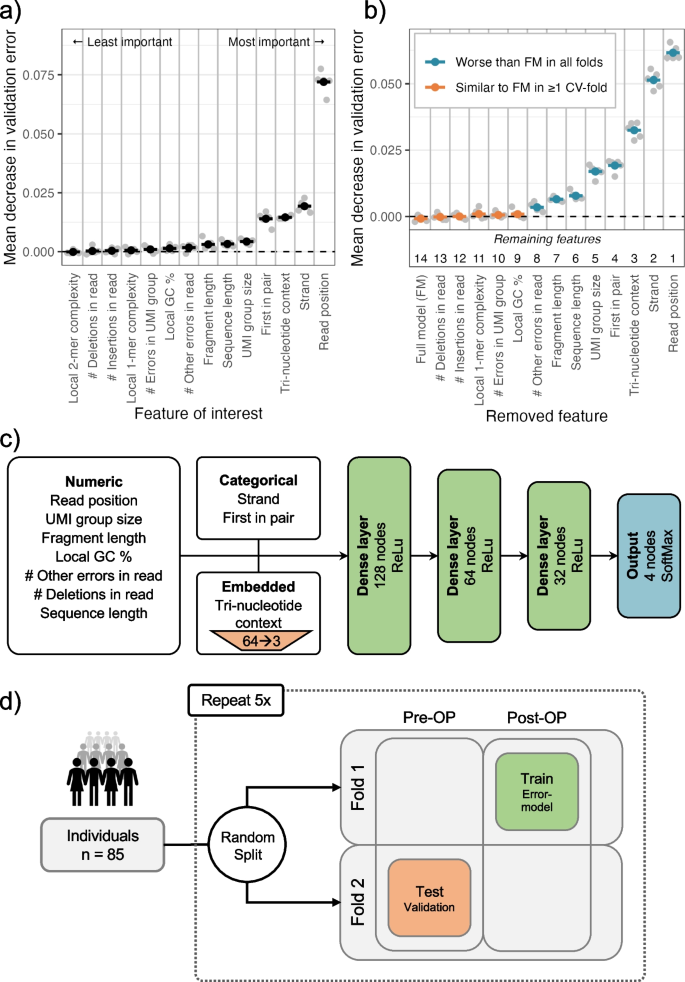
Circulating tumor DNA detection using next-generation sequencing (NGS) data of plasma DNA is promising for cancer identification and characterization. However, the tumor signal in the blood is often low and difficult to distinguish from errors. We present DREAMS (Deep Read-level Modelling of Sequencing-errors) for estimating error rates of individual read positions. Using DREAMS, we develop statistical methods for variant calling (DREAMS-vc) and cancer detection (DREAMS-cc). For evaluation, we generate deep targeted NGS data of matching tumor and plasma DNA from 85 colorectal cancer patients. The DREAMS approach performs better than state-of-the-art methods for variant calling and cancer detection.

Additional file 1 of DREAMS: deep read-level error model for

PDF) DREAMS: deep read-level error model for sequencing data

Integration of intra-sample contextual error modeling for improved
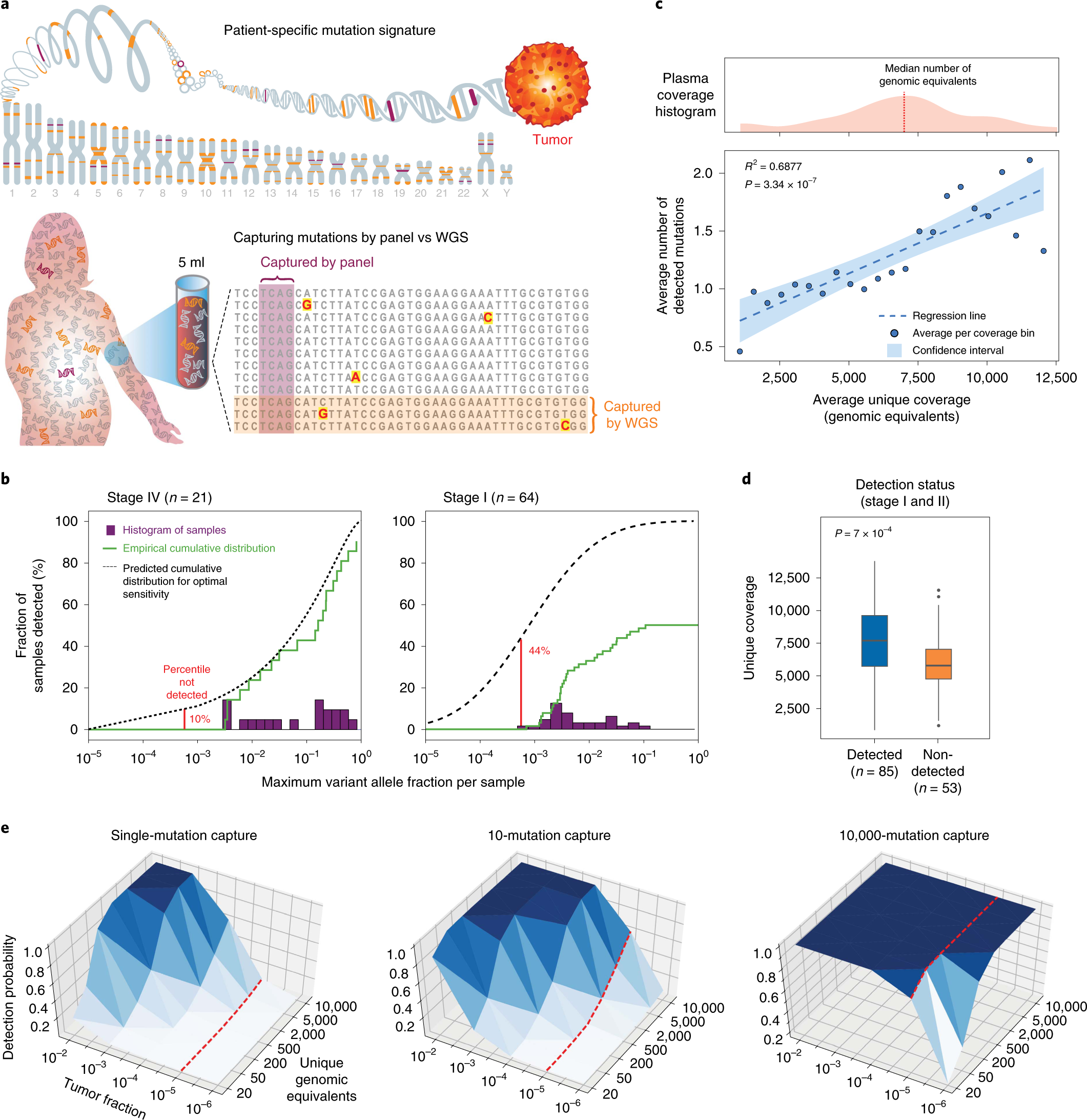
Genome-wide cell-free DNA mutational integration enables ultra
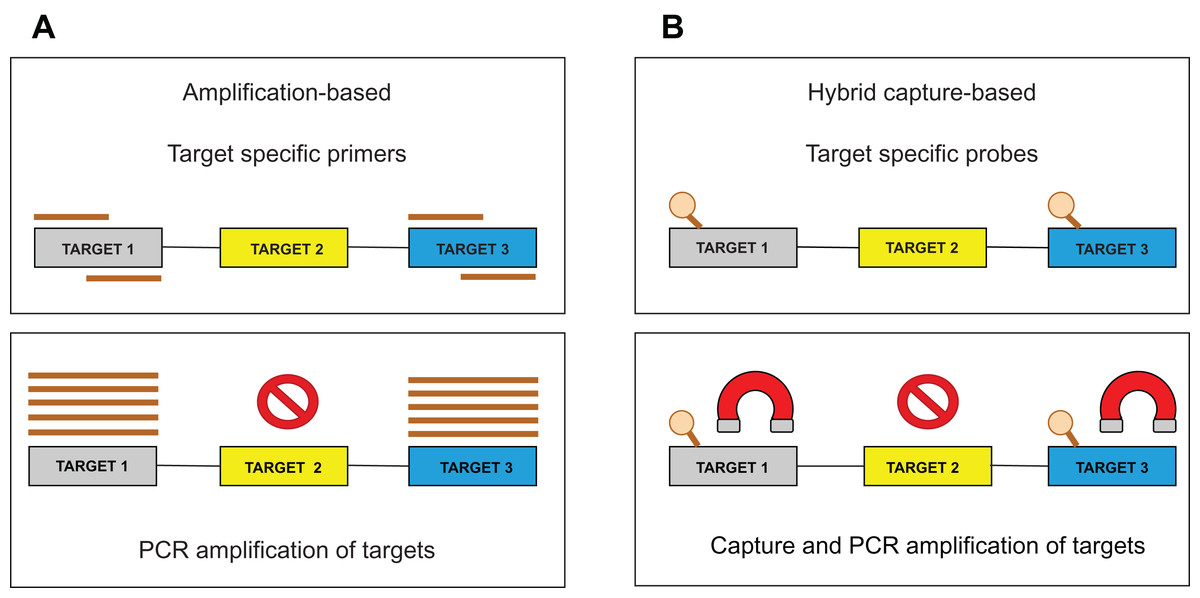
Bioinformatic strategies for the analysis of genomic aberrations
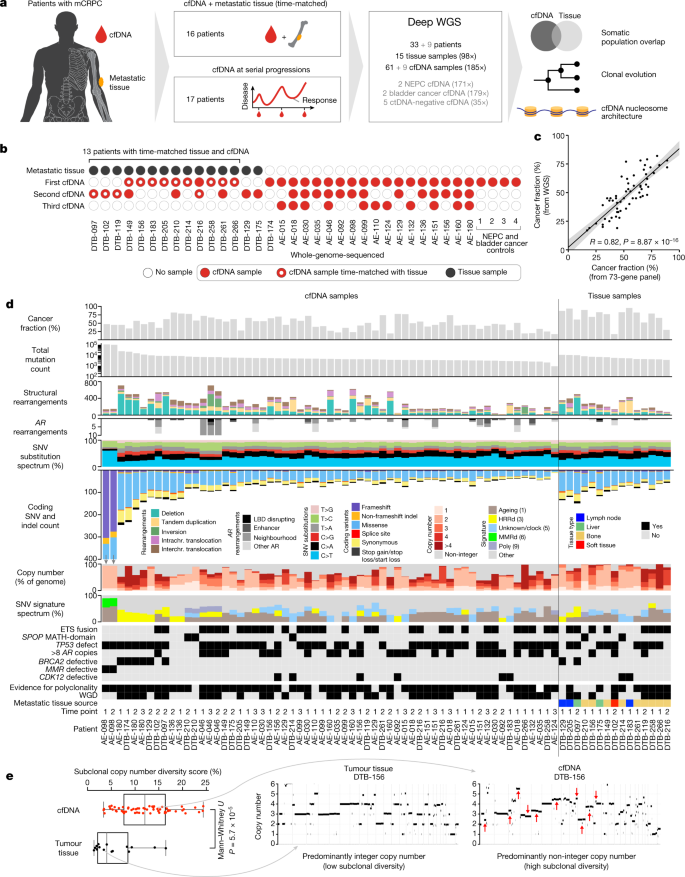
Deep whole-genome ctDNA chronology of treatment-resistant prostate

DREAMS: Deep Read-level Error Model for Sequencing data applied to
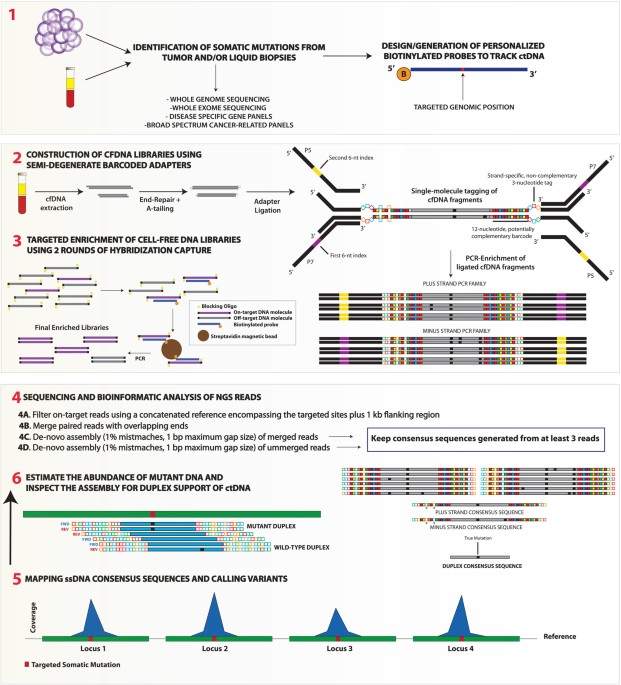
Targeted error-suppressed quantification of circulating tumor DNA

Analytical validation of an error-corrected ultra-sensitive ctDNA
Recomendado para você
-
 The Based God - SCP Foundation30 julho 2024
The Based God - SCP Foundation30 julho 2024 -
![7143-J SCP Foundation [RUS] Amino](http://pm1.aminoapps.com/7410/73c21043b027e12185482bb12bd40b31d49ade06r1-731-685v2_uhq.jpg) 7143-J SCP Foundation [RUS] Amino30 julho 2024
7143-J SCP Foundation [RUS] Amino30 julho 2024 -
scp #scpshowerthoughts #minecraft #minecraftparkour #scp173 #scpsecre30 julho 2024
-
 S I M P SCP Foundation (RP) Amino30 julho 2024
S I M P SCP Foundation (RP) Amino30 julho 2024 -
MisterMetokur - Welp the SCP wiki sanitized the doorknob30 julho 2024
-
 Emolog by Kyungmoo Min30 julho 2024
Emolog by Kyungmoo Min30 julho 2024 -
 Journal of Comparative Neurology, Systems Neuroscience Journal30 julho 2024
Journal of Comparative Neurology, Systems Neuroscience Journal30 julho 2024 -
parse_in/data.json at master · squiidz/parse_in · GitHub30 julho 2024
-
 Transgenic Mice Expressing Green Fluorescent Protein under the30 julho 2024
Transgenic Mice Expressing Green Fluorescent Protein under the30 julho 2024 -
 US20190218569A1 - Isolated polynucleotides and polypeptides and30 julho 2024
US20190218569A1 - Isolated polynucleotides and polypeptides and30 julho 2024
você pode gostar
-
 Análise: Pokémon Sword/Shield (Switch) traz a oitava geração dos monstrinhos mais queridos - Nintendo Blast30 julho 2024
Análise: Pokémon Sword/Shield (Switch) traz a oitava geração dos monstrinhos mais queridos - Nintendo Blast30 julho 2024 -
 Vale-Tudo Epic Developer Community30 julho 2024
Vale-Tudo Epic Developer Community30 julho 2024 -
 The Legend of Zelda™: Ocarina of Time™ Title Theme30 julho 2024
The Legend of Zelda™: Ocarina of Time™ Title Theme30 julho 2024 -
 Zombies.io for Android - Free App Download30 julho 2024
Zombies.io for Android - Free App Download30 julho 2024 -
 Stream Takumi Fujiwara Listen to Initial D First Stage: EP 2430 julho 2024
Stream Takumi Fujiwara Listen to Initial D First Stage: EP 2430 julho 2024 -
 Throne and Liberty: NCSoft anuncia data para pré-download e30 julho 2024
Throne and Liberty: NCSoft anuncia data para pré-download e30 julho 2024 -
 Rebel Moon - Parte 1: A Menina do Fogo (Filme), Trailer, Sinopse e30 julho 2024
Rebel Moon - Parte 1: A Menina do Fogo (Filme), Trailer, Sinopse e30 julho 2024 -
 Baixe Proton Bus Simulator Urbano no PC30 julho 2024
Baixe Proton Bus Simulator Urbano no PC30 julho 2024 -
_cb1b9562-57ea-45fd-8347-b7d06fe8d5f9.png) SÃO PAULO FC x EC BAHIA é na Total Acesso.30 julho 2024
SÃO PAULO FC x EC BAHIA é na Total Acesso.30 julho 2024 -
 Watch Chainsaw Man Online Free30 julho 2024
Watch Chainsaw Man Online Free30 julho 2024

